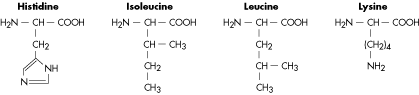|

CONSTANTS & EQUATIONS

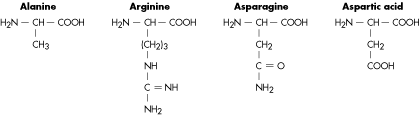
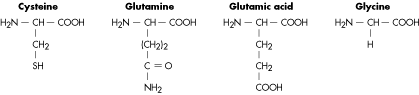
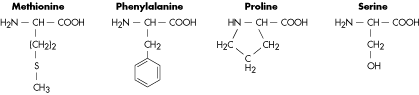
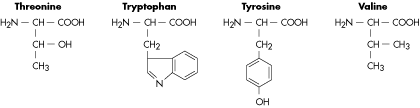
Molecular Structure of Amino Acids
 |
 |
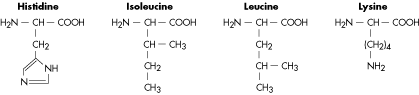 |
 |
 |
 [Return to top of page]
[Return to top of page]
The Genetic Code
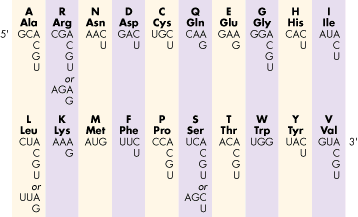 [Return to top of page]
[Return to top of page]
Nucleic Acid Data
1 µg of 1000 bp DNA = 1,52 pmol = 9.1 x 1011
molecules
1 µg of pUC18/19 DNA (2686 bp) = .057 pmol = 3.4 x 1011
molecules
1 µg of pBR322 DNA (4361 bp) = 0.35 pmol = 2.1 x 1011
molecules
1 µg of M13mp18/19 DNA (7250 bp) = 0.21 pmol = 1.3 x 1011
molecules
1 µg of l DNA (48502 bp) = 0.03 pmol 1.8 x 1010
molecules
1 pmol of 1000 bp DNA = 0.66 µg
1 pmol of pUC18/19 DNA (2686 bp) = 1.77 µg
1 pmol of l DNA (48502 bp) = 32.01 µg
[Return to top of page]
Spectrophotometric Conversions
1 A 260 unit of double-stranded DNA = 50 µg/ml
1 A 260 unit of single-stranded DNA = 33 µg/ml
1 A 260 unit of single-stranded RNA = 40 µg/ml
[Return to top of page]
Picomole Ends per Microgram of Double-Stranded Linear DNA
(2 x 106)/(660 x Number of bases) = pmol ends/µg double-stranded DNA
[Return to top of page]
Tm Factors Influencing
the Tm of Nucleic Acid Hybrids2
| Factor |
Influence on Tm |
| Ionic strength |
Tm increses 16.6° C for each onefold increase in monovalent
cations, between 0.01 and 0.40M NaCl |
Base
composition |
AT base pairs are less stable than GC base pairs in aqueous
solutions containing NaCI. The difference is negligible in tetramenthyl
ammoniuum chloride. |
Destablizing
agents |
Each 1% formamide reduces the Tm by about 0.6°C, 6M
urea reduces the Tm by about 30°C. |
Mismatched
base pairs |
The Tm is reduced by 1°C for each 1% of mismatching. |
| Duplex length |
Negligble effect with probes > 500 bp. |
[Return to top of page]
Equations for Calculating Tm
| System |
Equationa |
| DNA-DNA hybridsb |
Tm = 81.5°C + 16.6(logM) + 0.41(%GC) - .61(%form) -500/bp |
| DNA-RNA hybridsb |
Tm = 79.8°C + 18.5(logM) + 0.58(%GC) + 11.8(%GC)2
- 0.50(%form) - 820/L |
| RNA-RNA hybridsb |
Tm = 79.8°C + 18.5(logM) + 0.58(%GC) + 11.8(%GC)2
- 0.35(%form) - 820/L |
| Oligonucleotide probes |
Tm = 2(# AT bp) + 4(#GC bp) |
a M, molarity of monovalent cations (Na+
concentration, % GC, percentage of G and C nucleotides in the DNA, % form,
percentage of Formamide in the hybridization solutions; L, length of the
duplex in base pairs.
b These equations hold for Na+
concentrations between 0.01 and 0.40M and %GC values of 30-75%.
[Return to top of page]
References
1. Brown, T.A. Molecular Biology LabFax, Bios Scientific Publishers, Academic Press, 1991.
2. Dyson, N.J. (1991) In Essential Molecular Biology: A Practical
Approach. T.A. Brown (ed) Vol. 2 Oxford University Press, Oxford, in press.
3. Meinkoth, J. and Wahl G. (1984) Anal. Biochem., 138:267.
4.Casey, J. and Davidson, N. (1977) Nucleic Acids Res., 4:1539.
5. Bodkin, D.K. and Knudson, D.L. (1985) J. Virol. Methods, 10:45.
6. Wallace, R.B., et al. (1979) Nucleic Acids Res., 6:3543.
[Return to top of page]
|
|
|